library(tidyr)
library(dplyr)
library(purrr)
library(stringr)
library(forcats)
library(ggplot2)
library(scales)
library(patchwork)
library(janitor)
library(effsize)
library(metafor)
format_p_apa <- function(p, threshold = 0.001) {
ifelse(
p < threshold,
paste0("p < ", sub("^0\\.", ".", format(threshold, nsmall = 3))),
paste0("p = ", sub("^0\\.", ".", format(round(p, 3), nsmall = 3)))
)
}15 The differences between significant and non-significant is not itself significant ✎ Very rough draft
15.1 Dependencies
15.2 Simulation functions
# functions for simulation
generate_data <- function(n_per_condition,
population_cohens_d) {
data_control <-
tibble(condition = "control",
score = rnorm(n = n_per_condition, mean = 0, sd = 1))
data_intervention <-
tibble(condition = "intervention",
score = rnorm(n = n_per_condition, mean = population_cohens_d, sd = 1))
data_combined <- bind_rows(data_control,
data_intervention) |>
mutate(condition = fct_relevel(condition, "intervention", "control"))
return(data_combined)
}
analyze <- function(data, study) {
res_t_test <- t.test(formula = score ~ condition,
data = data,
var.equal = TRUE,
alternative = "two.sided")
res_cohens_d <- effsize::cohen.d(formula = score ~ condition,
data = data,
pooled = TRUE)
res <- tibble(study = study,
p = res_t_test$p.value,
cohens_d = res_cohens_d$estimate,
cohens_d_ci_lower = res_cohens_d$conf.int[1],
cohens_d_ci_upper = res_cohens_d$conf.int[2],
cohens_d_se = sqrt(res_cohens_d$var))
return(res)
}
meta_analyze <- function(results_study_1, results_study_2) {
dat <- bind_rows(results_study_1,
results_study_2)
fit <- rma(data = dat,
yi = cohens_d,
sei = cohens_d_se,
mods = ~ study,
method = "FE")
res <- tibble(cohens_d = fit$b[2],
cohens_d_ci_lower = fit$ci.lb[2],
cohens_d_ci_upper = fit$ci.ub[2],
#df = fit$QMdf[1],
#Q = fit$QM,
p = fit$QMp) # p of moderation
#mutate(formatted = paste0("Q(", df, ") = ", round_half_up(Q, 3), ", ", format_p_apa(p)))
return(res)
}15.3 Different population effect, large N
# set seed
set.seed(42)
# simulation parameters
experiment_parameters <- expand_grid(
study_1_n_per_condition = 300,
study_1_population_cohens_d = 0.5,
study_2_n_per_condition = 300,
study_2_population_cohens_d = 0,
iteration = 1:1000
)
# run simulation
simulation <- experiment_parameters |>
mutate(generated_data_study_1 = pmap(list(n_per_condition = study_1_n_per_condition,
population_cohens_d = study_1_population_cohens_d),
generate_data)) |>
mutate(generated_data_study_2 = pmap(list(n_per_condition = study_2_n_per_condition,
population_cohens_d = study_2_population_cohens_d),
generate_data)) |>
mutate(results_study1 = pmap(list(data = generated_data_study_1),
analyze,
study = "study 1")) |>
mutate(results_study2 = pmap(list(data = generated_data_study_2),
analyze,
study = "study 2")) |>
mutate(results_meta = pmap(list(results_study_1 = results_study1,
results_study_2 = results_study2),
meta_analyze)) |>
# wrangle results
unnest(results_study1, names_sep = "_") |>
unnest(results_study2, names_sep = "_") |>
unnest(results_meta, names_sep = "_") |>
select(-generated_data_study_1, -generated_data_study_2) |>
# this is a complex pivot, don't worry if you don't immediately understand it
pivot_longer(
cols = c(
results_study1_p,
results_study1_cohens_d,
results_study1_cohens_d_ci_lower,
results_study1_cohens_d_ci_upper,
results_study1_cohens_d_se,
results_study2_p,
results_study2_cohens_d,
results_study2_cohens_d_se,
results_study2_cohens_d_ci_lower,
results_study2_cohens_d_ci_upper,
results_meta_p,
results_meta_cohens_d,
# results_meta_cohens_d_se, # not created
results_meta_cohens_d_ci_lower,
results_meta_cohens_d_ci_upper
),
names_to = c("source", "metric"),
names_pattern = "results_(study\\d+|meta)_(.+)",
values_to = "value"
) |>
pivot_wider(
names_from = metric,
values_from = value
)
simulation_summary <- simulation |>
group_by(study_1_n_per_condition,
study_1_population_cohens_d,
study_2_n_per_condition,
study_2_population_cohens_d,
source) |>
summarize(proportion_significant = mean(p < .05),
mean_cohens_d = mean(cohens_d),
mean_cohens_d_ci_lower = mean(cohens_d_ci_lower),
mean_cohens_d_ci_upper = mean(cohens_d_ci_upper),
.groups = "drop") |>
mutate(source = str_replace(source, "meta", "difference")) |>
mutate(source = fct_relevel(source, "study1", "study2", "difference"))
p_effect_size <- ggplot(simulation_summary, aes(source, mean_cohens_d)) +
geom_hline(yintercept = 0, linetype = "dotted") +
geom_linerange(aes(ymin = mean_cohens_d_ci_lower, ymax = mean_cohens_d_ci_upper)) +
geom_point() +
scale_y_continuous(breaks = breaks_pretty(n = 10),
name = "Mean Cohen's d") +
xlab("") +
theme_linedraw() +
theme(panel.grid.minor = element_blank())
p_proportion_significant <- ggplot(simulation_summary, aes(source, proportion_significant)) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
geom_hline(yintercept = 0.80, linetype = "dashed") +
geom_bar(stat = "identity") +
scale_y_continuous(breaks = breaks_pretty(n = 10), limits = c(0, 1), name = "Proportion of significant p values") +
xlab("") +
# geom_text(aes(x = "study1", y = 1.1, label = paste("Population\nCohen's d =", study_1_population_cohens_d))) +
# geom_text(aes(x = "study2", y = 1.1, label = paste("Population\nCohen's d =", study_2_population_cohens_d))) +
# geom_text(aes(x = "difference", y = 1.1, label = paste("Population\nCohen's d =", study_2_population_cohens_d - study_1_population_cohens_d))) +
theme_linedraw() +
theme(panel.grid.minor = element_blank())
p_effect_size + p_proportion_significant + plot_layout(ncol = 2)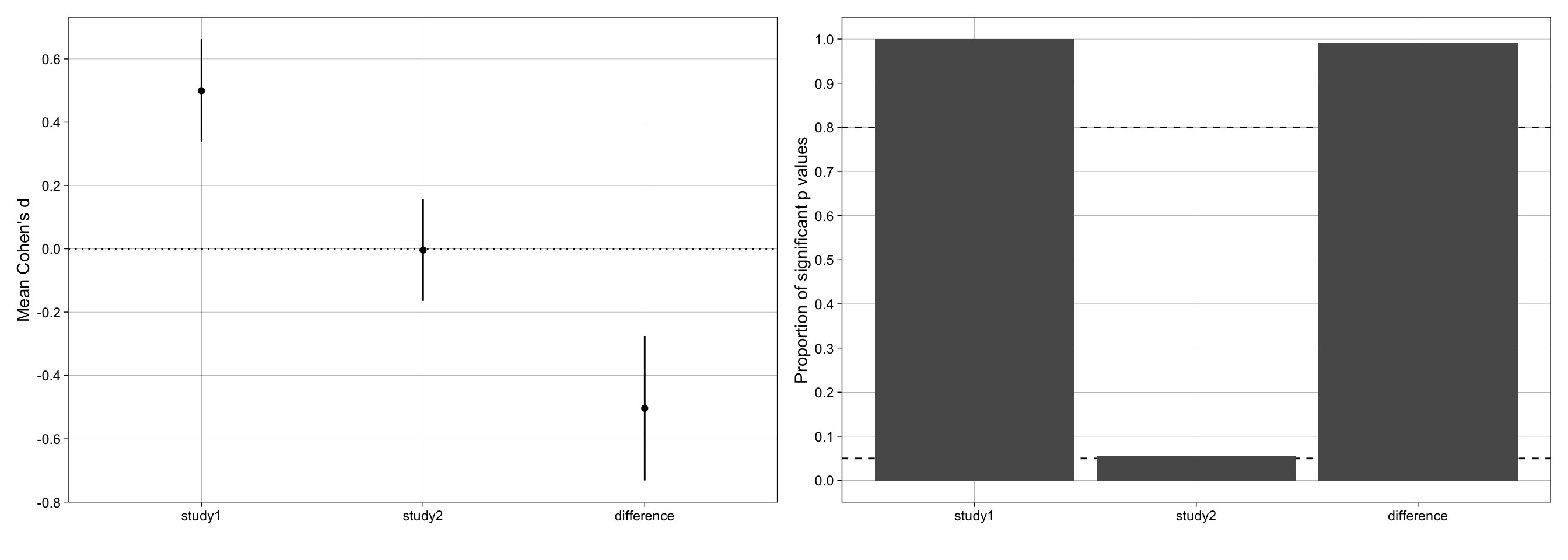
15.4 Different population effect, small N
# set seed
set.seed(42)
# simulation parameters
experiment_parameters <- expand_grid(
study_1_n_per_condition = 40,
study_1_population_cohens_d = 0.5,
study_2_n_per_condition = 40,
study_2_population_cohens_d = 0,
iteration = 1:1000
)
# run simulation
simulation <- experiment_parameters |>
mutate(generated_data_study_1 = pmap(list(n_per_condition = study_1_n_per_condition,
population_cohens_d = study_1_population_cohens_d),
generate_data)) |>
mutate(generated_data_study_2 = pmap(list(n_per_condition = study_2_n_per_condition,
population_cohens_d = study_2_population_cohens_d),
generate_data)) |>
mutate(results_study1 = pmap(list(data = generated_data_study_1),
analyze,
study = "study 1")) |>
mutate(results_study2 = pmap(list(data = generated_data_study_2),
analyze,
study = "study 2")) |>
mutate(results_meta = pmap(list(results_study_1 = results_study1,
results_study_2 = results_study2),
meta_analyze)) |>
# wrangle results
unnest(results_study1, names_sep = "_") |>
unnest(results_study2, names_sep = "_") |>
unnest(results_meta, names_sep = "_") |>
select(-generated_data_study_1, -generated_data_study_2) |>
# this is a complex pivot, don't worry if you don't immediately understand it
pivot_longer(
cols = c(
results_study1_p,
results_study1_cohens_d,
results_study1_cohens_d_ci_lower,
results_study1_cohens_d_ci_upper,
results_study1_cohens_d_se,
results_study2_p,
results_study2_cohens_d,
results_study2_cohens_d_se,
results_study2_cohens_d_ci_lower,
results_study2_cohens_d_ci_upper,
results_meta_p,
results_meta_cohens_d,
# results_meta_cohens_d_se, # not created
results_meta_cohens_d_ci_lower,
results_meta_cohens_d_ci_upper
),
names_to = c("source", "metric"),
names_pattern = "results_(study\\d+|meta)_(.+)",
values_to = "value"
) |>
pivot_wider(
names_from = metric,
values_from = value
)
simulation_summary <- simulation |>
group_by(study_1_n_per_condition,
study_1_population_cohens_d,
study_2_n_per_condition,
study_2_population_cohens_d,
source) |>
summarize(proportion_significant = mean(p < .05),
mean_cohens_d = mean(cohens_d),
mean_cohens_d_ci_lower = mean(cohens_d_ci_lower),
mean_cohens_d_ci_upper = mean(cohens_d_ci_upper),
.groups = "drop") |>
mutate(source = str_replace(source, "meta", "difference")) |>
mutate(source = fct_relevel(source, "study1", "study2", "difference"))
p_effect_size <- ggplot(simulation_summary, aes(source, mean_cohens_d)) +
geom_hline(yintercept = 0, linetype = "dotted") +
geom_linerange(aes(ymin = mean_cohens_d_ci_lower, ymax = mean_cohens_d_ci_upper)) +
geom_point() +
scale_y_continuous(breaks = breaks_pretty(n = 10),
name = "Mean Cohen's d") +
xlab("") +
theme_linedraw() +
theme(panel.grid.minor = element_blank())
p_proportion_significant <- ggplot(simulation_summary, aes(source, proportion_significant)) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
geom_hline(yintercept = 0.80, linetype = "dashed") +
geom_bar(stat = "identity") +
scale_y_continuous(breaks = breaks_pretty(n = 10), limits = c(0, 1), name = "Proportion of significant p values") +
xlab("") +
# geom_text(aes(x = "study1", y = 1.1, label = paste("Population\nCohen's d =", study_1_population_cohens_d))) +
# geom_text(aes(x = "study2", y = 1.1, label = paste("Population\nCohen's d =", study_2_population_cohens_d))) +
# geom_text(aes(x = "difference", y = 1.1, label = paste("Population\nCohen's d =", study_2_population_cohens_d - study_1_population_cohens_d))) +
theme_linedraw() +
theme(panel.grid.minor = element_blank())
p_effect_size + p_proportion_significant + plot_layout(ncol = 2)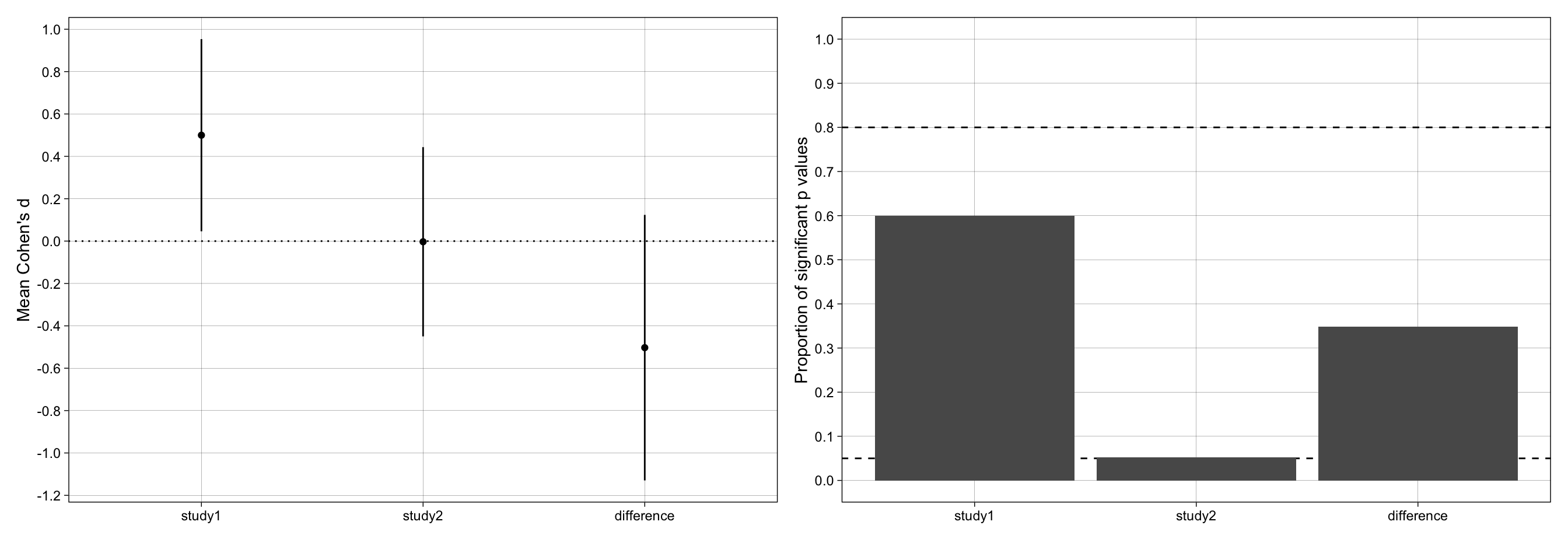
15.5 Different population effect, small original study and large “replication”
NB not really a replication as the second study has a different effect size, but its a very rough simulation assuming p hacking took place in the original study and boosted the effect size
[needs work to be clearer]
# set seed
set.seed(42)
# simulation parameters
experiment_parameters <- expand_grid(
study_1_n_per_condition = 40,
study_1_population_cohens_d = 0.5,
study_2_n_per_condition = 200,
study_2_population_cohens_d = 0,
iteration = 1:1000
)
# run simulation
simulation <- experiment_parameters |>
mutate(generated_data_study_1 = pmap(list(n_per_condition = study_1_n_per_condition,
population_cohens_d = study_1_population_cohens_d),
generate_data)) |>
mutate(generated_data_study_2 = pmap(list(n_per_condition = study_2_n_per_condition,
population_cohens_d = study_2_population_cohens_d),
generate_data)) |>
mutate(results_study1 = pmap(list(data = generated_data_study_1),
analyze,
study = "study 1")) |>
mutate(results_study2 = pmap(list(data = generated_data_study_2),
analyze,
study = "study 2")) |>
mutate(results_meta = pmap(list(results_study_1 = results_study1,
results_study_2 = results_study2),
meta_analyze)) |>
# wrangle results
unnest(results_study1, names_sep = "_") |>
unnest(results_study2, names_sep = "_") |>
unnest(results_meta, names_sep = "_") |>
select(-generated_data_study_1, -generated_data_study_2) |>
# this is a complex pivot, don't worry if you don't immediately understand it
pivot_longer(
cols = c(
results_study1_p,
results_study1_cohens_d,
results_study1_cohens_d_ci_lower,
results_study1_cohens_d_ci_upper,
results_study1_cohens_d_se,
results_study2_p,
results_study2_cohens_d,
results_study2_cohens_d_se,
results_study2_cohens_d_ci_lower,
results_study2_cohens_d_ci_upper,
results_meta_p,
results_meta_cohens_d,
# results_meta_cohens_d_se, # not created
results_meta_cohens_d_ci_lower,
results_meta_cohens_d_ci_upper
),
names_to = c("source", "metric"),
names_pattern = "results_(study\\d+|meta)_(.+)",
values_to = "value"
) |>
pivot_wider(
names_from = metric,
values_from = value
)
simulation_summary <- simulation |>
group_by(study_1_n_per_condition,
study_1_population_cohens_d,
study_2_n_per_condition,
study_2_population_cohens_d,
source) |>
summarize(proportion_significant = mean(p < .05),
mean_cohens_d = mean(cohens_d),
mean_cohens_d_ci_lower = mean(cohens_d_ci_lower),
mean_cohens_d_ci_upper = mean(cohens_d_ci_upper),
.groups = "drop") |>
mutate(source = str_replace(source, "meta", "difference")) |>
mutate(source = fct_relevel(source, "study1", "study2", "difference"))
p_effect_size <- ggplot(simulation_summary, aes(source, mean_cohens_d)) +
geom_hline(yintercept = 0, linetype = "dotted") +
geom_linerange(aes(ymin = mean_cohens_d_ci_lower, ymax = mean_cohens_d_ci_upper)) +
geom_point() +
scale_y_continuous(breaks = breaks_pretty(n = 10),
name = "Mean Cohen's d") +
xlab("") +
theme_linedraw() +
theme(panel.grid.minor = element_blank())
p_proportion_significant <- ggplot(simulation_summary, aes(source, proportion_significant)) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
geom_hline(yintercept = 0.80, linetype = "dashed") +
geom_bar(stat = "identity") +
scale_y_continuous(breaks = breaks_pretty(n = 10), limits = c(0, 1), name = "Proportion of significant p values") +
xlab("") +
# geom_text(aes(x = "study1", y = 1.1, label = paste("Population\nCohen's d =", study_1_population_cohens_d))) +
# geom_text(aes(x = "study2", y = 1.1, label = paste("Population\nCohen's d =", study_2_population_cohens_d))) +
# geom_text(aes(x = "difference", y = 1.1, label = paste("Population\nCohen's d =", study_2_population_cohens_d - study_1_population_cohens_d))) +
theme_linedraw() +
theme(panel.grid.minor = element_blank())
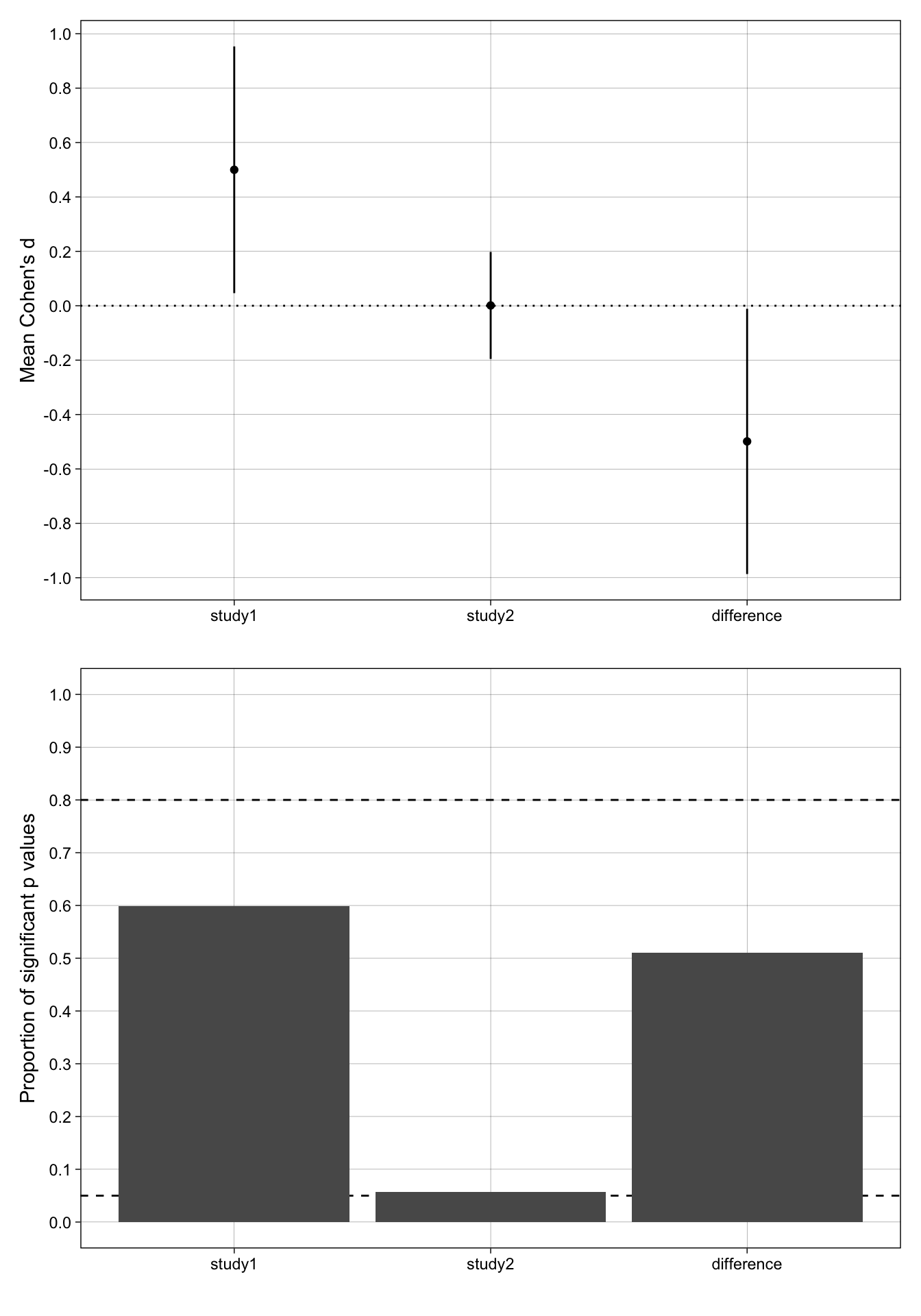
p_effect_size + p_proportion_significant + plot_layout(ncol = 1)